Navigation auf uzh.ch
Navigation auf uzh.ch

Heavy metals such as cadmium (Cd) and lead (Pb) are important environmental pollutants with high toxicity to animals and plants. They inactivate or denature proteins by binding to sulfhydryl groups, causing cellular damage by displacing cofactors from a variety of proteins including transcription factors and enzymes, and by indirectly generating reactive oxygen species. Heavy metal detoxification mechanisms in plants and other organisms employ chelating compounds such as metallothioneins and phytochelatins, Cys-rich membrane proteins and heavy metal transporters. Different types of transporters either directly energized by ATP such as P-type ATPases and ABC transporters or energized by the proton motive force, such as NRAMPs or ZIPs, have been shown to be involved in heavy metal resistance.
We are interested to elucidate the role of ABC transporters in heavy metal transport and resistance. ABC transporters constitute one of the largest families of proteins in living organisms ranging from bacteria to humans. ABC proteins are involved in the transport of a wide variety of substances, including ions, carbohydrates, lipids, xenobiotics, antibiotics, drugs, and heavy metals. Among the many plant ABC transporters AtATM3, AtPDR12 and AtPDR8 play an important role in heavy metal resistance. Our goal is to elucidate the transport mechanism of these transporters and to find new ABC transporters involved in heavy metal transport. An applied aspect of this project is the use of our knowledge to create plants, which are more heavy metal tolerant, take up more heavy metals and can be used for phytoremediation. Phytoremediation is an environmental friendly strategy to clean up heavy metal contaminated soils by plants extracting the heavy metals from the soil. Our knowledge will also be suited to find traits for breeding crops, which take up a minimum of heavy metals and hence reduce the amount of toxic metals entering the food chain.
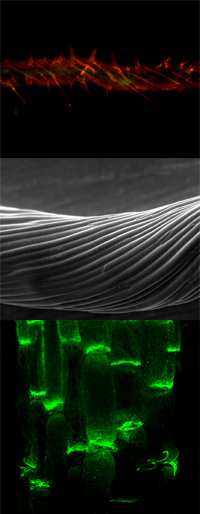
Many aspects of plant physiology and development are dependent on the polar transport of the plant signaling compound auxin (IAA). However, cellular efflux is the rate-limiting step of this unique cell-to-cell movement of IAA, called PAT.
Our work concentrates on deciphering the individual roles and functional interaction of catalytic and regulatory components of auxin transport complexes and their involvement in auxin-mediated plant physiology and development.
Recent key findings
We have identified ABCB/PGP/MDR (P-GLYCOPROTEIN, MULTIDRUG-RESISTANCE) proteins as novel, primary active (ATP-dependent) auxin pumps showing auxin-related, developmental (but no organogenetic) loss-of-function phenotypes. ABCB1 and ABCB19 are, unlike their mammalian orthologs, specific to auxins and, despite their mostly apolar locations, function in long-range auxin transport. For ABCB4 an inward transport direction has been demonstrated, not yet found for non-plant ABCBs.
ABCB- and PIN-mediated auxin efflux can function independently, and play identical cellular but separate developmental roles. Moreover, ABCBs and PINs are able to interactively and coordinately transport auxin. In interacting cells, the current model is that multilaterally ABCBs minimize apoplastic reflux while polar ABCB-PIN interactions provide vectorial auxin streams.
The immunophilin-like FKBP42, TWD1 (TWISTED DWARF 1), has been characterized as central regulator of ABCB- and ABCC-mediated transport by means of protein-protein interaction. Positive regulation of ABCB1,19-mediated auxin transport accounts for overlapping phenotypes between twd1 and abcb1,19. The diagnostic auxin transport inhibitor, NPA, disrupts TWD1-ABCB1 interaction by binding to ABCB1 and TWD1. This leads to disruption of ABCB1 activity, suggesting that TWD1 and ABCB1 represent key components of the NPA-sensitive efflux complex.
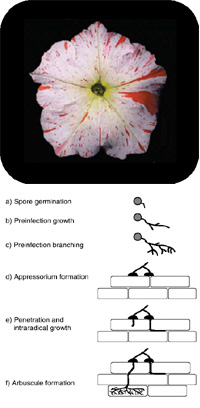
Arbuscular Mycorrhiza (AM) comprises the mutual symbiotic relationship between soil born fungi of the order Glomales and an estimated 80% of all vascular plants. The establishment of the symbiosis, from spore germination of the biotrophic AM fungus to arbuscule formation in host root cortex cells, depends on a well-balanced exchange of signaling compounds both of stimulatory and inhibitory nature. In each of the symbiotic steps a multitude of secondary compounds, such as simple phenolic, flavonoid and terpenoid compounds, have been demonstrated to be differentially produced and/or excreted. Loss of function mutants in terpenoid biosynthesis exhibit a severe deficiency in mycorrhization, indicating a functional role of these metabolites.
However, up to date no studies have investigated the role of secondary metabolite transporters, such as ABC transporters, in this context. The subfamily of Pleiotropic Drug Resistance (PDR) proteins appears to be the most interesting. PDRs, with 15 members in Arabidopsis, are exclusively found in the plant and fungal kingdoms. In the former, they are often highly expressed in roots, seem to predominantly reside in the plasma membrane, have been demonstrated to transport terpenoids and are implicated in biotic and abiotic stress response.
In order to gain further insight into the putative role of PDRs and their substrates in AM, we attempt to identify mycorrhiza-responsive members of the PDR subfamily in the transposon-harboring plant Petunia hybridia W138. We then aim to retrieve W138 lines with transposon insertions in our genes of interest and analyze homozygous mutants for phenotypic alterations in regard to mycorrhization and root exudation.
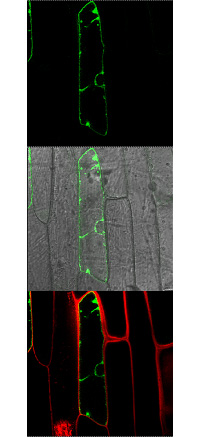
In mature plant cells the vacuole is by far the largest organelle and occupies up to 90% of the cell volume. The vacuole serves as a temporary store for a large number of metabolites and ions, such as sugars, organic acids, amino acids, potassium, phosphate and nitrate just to mention a few. Since for an optimal function of the metabolism cytosolic concentrations have to be maintained constant for many metabolites, vacuolar import, storage and export are the prerequisite for the cytosolic homeostasis of most compounds. The identification and characterization of vacuolar membrane proteins was much slower compared to the plasma membrane located proteins, mostly because no appropriate yeast mutants were available allowing complementation studies. However, in the last few years considerable progress has been made and a large number of vacuolar membrane proteins have been identified and characterized. This progress was due to the complete sequencing of the Arabidopsis genome, the comparison of Arabidopsis with already known sequences and the generation of knockout lines for nearly all Arabidopsis genes.
The goal of this project is to identify and characterize vacuolar channels and transporters. To reach this goal we are using different approaches such as vacuolar proteomics, phosphoproteomics and quantitative proteomics, molecular biology, reverse genetics, physiology and electrophysiology. We also aim to integrate metabolomics in the near future. Our knowledge will help to understand better how cellular metabolism is regulated.
(Project leader: Dr. Diana Santelia)
We are interested in understanding the molecular mechanisms of carbon partitioning at different stages of water deficit in the model specie Arabidopsis thaliana.
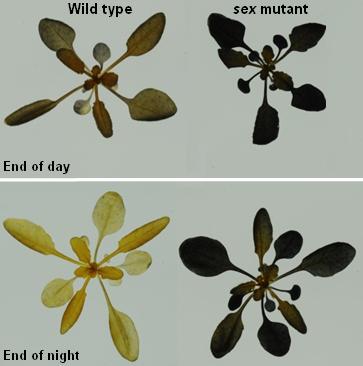

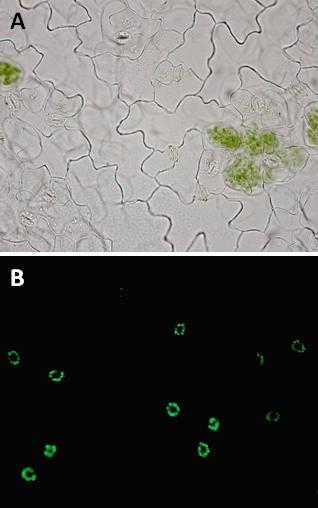
In the case of mild water stress, the partial closure of stomata can be sufficient to sustain plant performance, thus making guard cells a putative target to engineer drought avoidance in plants with minimal alterations of yield. The success in engineering stomatal function depends upon a detailed molecular understanding of guard cell osmoregulation processes. In guard cells, the opening of stomata at the onset of light is driven by massive uptake of K+ ions from the apoplast, accompanied by malate2- synthesis in the cytosol. In the current view, storage and mobilization of starch in guard cells follow an opposite rhythm with respect to mesophyll cells. Namely, starch is present in darkness in almost all guard cells and is degraded during the first hours of light, presumably to provide carbon precursors needed for malate synthesis or sucrose accumulation. While the metabolism of K+ and malate2- has been extensively studied, the precise role of starch and sucrose in guard cells remain largely unknown, despite the importance that has been recognize to these compounds in the regulation of stomatal movement.
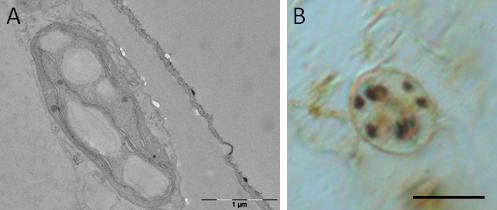
We employ a diverse array of molecular and biochemical approaches to investigate the cellular mechanisms of starch breakdown in Arabidopsis guard cells and to identify the key factors required to trigger starch degradation at the onset of light. We also investigate how the stress-induced reprogramming of carbon partitioning is affecting malate and/or sucrose metabolism in guard cells, and what consequences might have on stomatal behavior during water deficit.
In case of prolonged or more severe stress, one of the most important mechanisms of drought resistance is the accumulation of soluble sugars to avoid water loss and to protect membranes, enzymes and other structures against damage and denaturation. In most cases, the accumulation of non-reducing sugars (mainly sucrose, fructose and glucose) in response to dehydration stress begins after the cessation of photosynthesis. It is therefore a result of an alteration in carbon partitioning (most likely from starch to soluble sugars) and not a product of carbon assimilation. In this respect, starch breakdown in the early phases of acute stress can be considered as a short-term mechanism of survival for the sustainment of the central carbohydrate metabolism in the production of osmoprotectants and energy generation. The biochemical mechanisms that control and influence the flow of carbon upon stress have not been fully assessed or remain unknown. Moreover, an overview of the consequences of a severe water deficit on carbon partitioning in Arabidopsis thaliana is still missing.
To identify the precise metabolic origin of the increased sugar pool caused by water deficit and the way the stress-induced reprogramming of carbon metabolism is regulated, we perform 14CO2 “pulse-chase” experiments in Arabidopsis wild type plants and a set of starch/sucrose-related mutants exposed to different levels of water stress. We also employ forward genetics approaches to identify transcriptional and posttranslational regulatory networks controlling the rapid reprogramming of carbohydrate metabolism upon water stress perception in Arabidopsis.