Navigation auf uzh.ch
Navigation auf uzh.ch
Microbial pathogens are an integral part of earth ecosystem, and virtually all living organisms are challenged by pathogens at some stage of their life. The study of pathogens profited greatly from the improvements in sequencing techniques of the last two decades. The drop in the cost of sequencing, and the development of statistical and computational methods to analyze sequence data, opened the way to large scale population genomic analyses. For many human pathogens, population genomics and molecular epidemiology have become crucial tools for research, but also for surveillance, monitoring and diagnostic for public health.
It has long been recognized that understanding the population genetics of agricultural plant pathogens could contribute to improve disease control, and in the past decades much effort was placed to study the genetic structure of pathogen populations, their migration routes, host shifts and reproductive strategies However, because of reduced resources and technical challenges such as the large genome size of many plant pathogens, population genetics and molecular epidemiology of agricultural pathogens have not been characterized by the large scale real-time genomic approach that was so fruitful in the study of human infectious diseases.
Our current projects use large scale genomics and molecular epidemiology to study wheat powdery mildew (Blumeria graminis forma specialis tritici), a fungal pathogen of wheat.
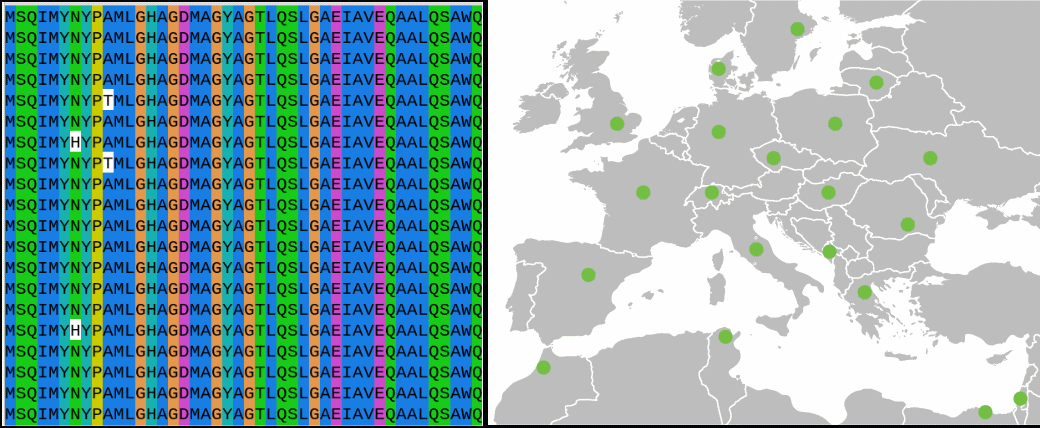
We are investigating the population genetics of wheat powdery mildew in Europe at different geographic scales, from a single field to the whole continent. Our goal is to understand the seasonal epidemics of wheat powdery mildew, and its recent evolutionary history. We try to characterize the extent of sexual reproduction in different populations, and track the spread of different genotypes over time to determine the spatiotemporal dynamics of the epidemics, such as the main directions of dispersal and the origin of the inoculum each year.