Navigation auf uzh.ch
Navigation auf uzh.ch

Brachypodium distachyon is a wild grass which occur around all the Mediterranean rim. B. distachyon has been developed as a powerful model for research on temperate grass species as it is closely related to major crop cereals and to some of the grasses used for biofuel production. Entirely sequenced, its small diploid genome (280Mb) is fully assembled into five chromosomes and has been exhaustively annotated. It represents therefore a prime system for research in population genomics and ecology.
See https://jgi.doe.gov/our-science/science-programs/plant-genomics/brachypodium/
Transposable elements (TEs) are mobile DNA sequences which have the capacity to replicate and move from one location to another in the genome. While TEs have long been seen as causing agents of genomic instability, it is now established that they can produce broad functional and beneficial changes, from the fine tuning of nearby gene expression to the alteration of gene function. They constitute therefore a driving force of evolution.
TEs activity is induced by biotic and abiotic stresses. Thus, distinct ecological environments acting on closely related lineages are expected to create intra-specific variability in TE patterns (genetic novelty). However, with the reliance of most research on plant of economical interest, examples enlightening the role of TEs on phenotypic variation are bias towards visible traits of agronomical interest. For these reasons, we still ignore how often and to what extent TEs contribute to adaptive evolution in plants. Using natural accessions of B. distachyon, our first project aims at better characterizing the role of TE polymorphisms on natural populations evolution and local adaptation.
We are thankful to Mhemmed Gandour, Abdelmonaim Homrani Bakali and Danka Petrovic for providing us with samples from Tunisia, Morocco and Montenegro.
We developed a new approach to map TE insertion polymorphisms using reference-aligned paired-end reads. If you are interested, check https://github.com/cstritt/detettore
Because insertions into coding or regulatory sequences can be highly deleterious, active TEs pose a considerable threat to genome integrity. Yet, host genomes have evolved safeguarding mechanisms enabling TE silencing. Typically, RNA-directed DNA Methylation (RdDM) epigenetically maintains TEs in a transcriptionally silent state via DNA and histone methylation. As repressive epigenetic states can spread to neighboring regions, TEs can influence the expression of neighboring genes Strikingly, the genome of B. distachyon harbor numerous TE insertions in the vicinity of genes (<2kb). Our second project aim at answering fascinating questions about the role of TE on epigenetic regulation: what is the genome-wide impact of these TE insertion polymorphisms on genome expression? Do novel TEs insertions modify local epigenetic states and spatial arrangements at target loci?
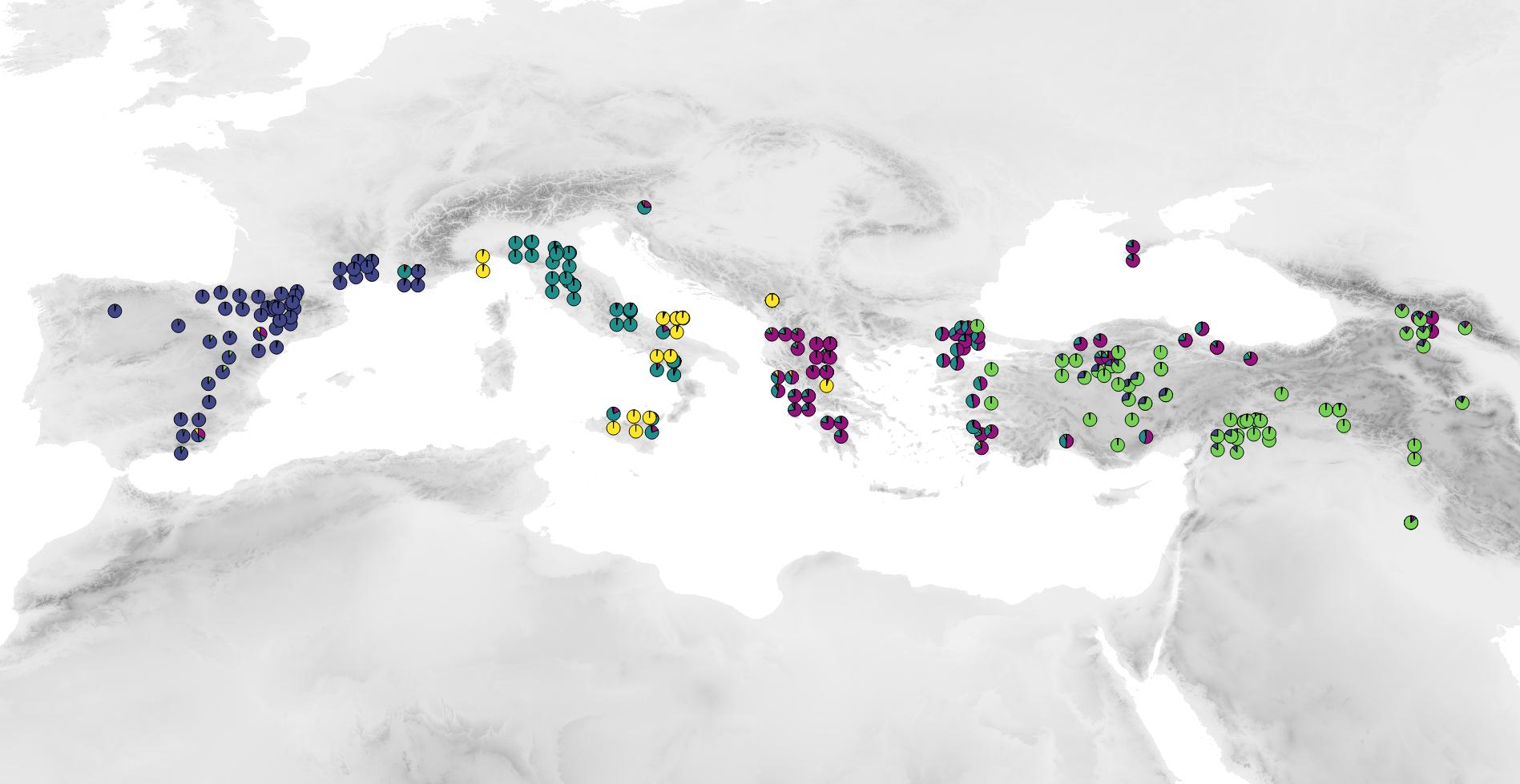
To understand how population adapt to their local environment also necessitate to better comprehend their demographic history as well as to characterise the selective pressure that act locally on them. Making use of the 143 accessions from Spain, France, Italy, Turkey and Iraq sequenced, we are currently investigating how bottlenecks and expansions shaped the distribution as well as the genetic diversity of B. distachyon. Our data show that all studied populations went through a bottleneck during the last glaciation but underwent expansion during that last 10,0000 years. Complementary analyses suggest that B. distachyon presents a similar history as A. thaliana: relict populations, i.e. populations that persisted in European refugees during the last glaciation spread over Europe when the climate warmed-up and nowadays co-occur with accessions which more recently recolonized Europe from North Africa or Middle-east. While this study will be extended to Grece, Montenegro and North- Africa in the near future, our analysis suggests that subsequent to the ending of the last glaciation and the colonization of Europe by Mediterranean plants, B. distachyon accessions locally adapted to newly colonised habitats. A genome-scan of positive selection lead on a subset of these accessions indeed revealed that selection acted recently on those population and showed that among other constraints, pathogens may constitute an important selection pressure in our system.